The mission of the UW Carbone Developmental Therapeutics (DT) Program is to improve cancer patient outcomes by discovering new targets, developing therapeutic agents and biomarkers, and translating this research into early phase clinical trials.
Research Areas
The overarching goal of the DT Program is to identify more effective therapies and strategies for the treatment of patients with advanced cancer. The DT Program houses much of UW Carbone’s drug and biomarker discovery and development work, as well as preclinical and clinical translational efforts.
Thematic Aim 1: Discover new molecular targets for cancer therapy. DT investigators are actively engaged in basic and translational research to identify new molecular targets for cancer therapy.
Thematic Aim 2: Develop new agents and biomarkers. New molecular targets will be translated by the timely development of therapeutic approaches. This includes isolation and characterization of new therapeutic agents and development of biomarkers and therapy-predictive assays for efficient translation of new agents and biomarkers.
Thematic Aim 3: Translate new therapies and biomarkers into clinical trials. New targets, agents, and biomarkers identified and developed within the DT Program (and other programs) are translated in proof-of-concept early phase clinical trials
Program Leaders
Christian Capitini
Dustin Deming
Beth Weaver
Developmental Therapeutics Membership
Select Recent Accomplishments
This is an accordion element with a series of buttons that open and close related content panels.
Biomarkers
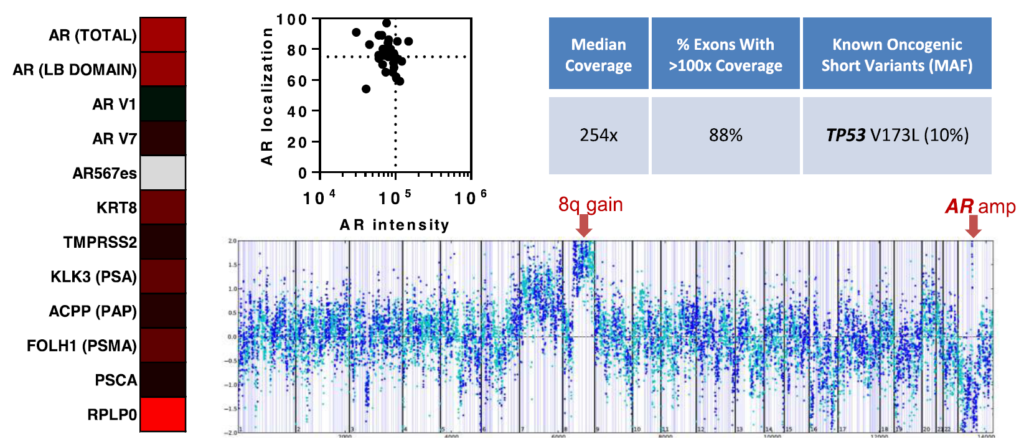
In addition to cell free DNA, circulating tumor cell (CTC) analysis permits frequent, minimally invasive sampling of tumor cells for molecular assessment. An example of Aim 2 research being applied to clinical trials is being conducted by Lang, a DOD Physician Research Training Award recipient, in collaboration with Beebe (TM). They have optimized a microfluidic technology known as the VERSA (Vertical Exclusion-based Rare Sample Analysis) that integrates capture and analysis of rare tumor cells for protein, gene expression, and genomic information. VERSA utilizes a unique separation principle that enables multiple purification processes in sequence (e.g., cell capture, extracellular staining, nucleic acid extraction) for sequential extraction of multiple analytes from a single sample.
Cancer Immunotherapy
DNA vaccines represent a cost-effective approach to induce antigen-specific immunity. The McNeel lab has advanced a DNA vaccine against human Prostatic Acid Phosphatase (PAP) and the Androgen-Receptor (AR) Ligand-Binding Domain (LBD) into clinical trials. With funding from the NIH/NCI and a DOD Prostate Cancer Research Program Clinical Trial Award, a Phase I and II dose- and schedule-evaluation trial was completed in patients with PSA-recurrent prostate cancer demonstrating that DNA immunization could safely elicit antigen-specific T-cell responses.
Quantitative Total Bone Imaging
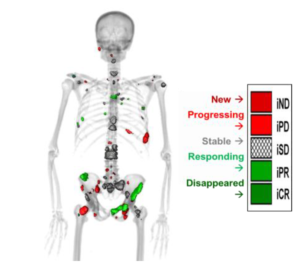
Quantitative Total Bone Imaging (QTBI) is a robust methodology developed by Jeraj and Liu to allow total and inter-lesional assessment of treatment response in osseous metastases. Their multi-center trial (UWCCC, Karmanos, and Rutgers) “A Positron Emission Tomography/Computed Tomography Bone Imaging Study in Patients Receiving Enzalutamide for Castration-Resistant Prostate Cancer” is ongoing. The purpose of this study is to prospectively evaluate the role of NaF PET/CT as a treatment response biomarker in mCRPC to bone.